By Nermina Lamadema, Postdoctoral Research Associate at King's College London
There is well over 200 different disease that come under the umbrella term cancer. Most of the cells in our body have a finite life span termed Hayflick limit which is a number of divisions after which a cell will commit to either die or enter a senescence which is equivalent to deep sleep. The mitotic cell cycle involves the duplication of chromosomes and other cellular components and their subsequent segregation into each of two daughter cells. The fidelity of the process, its timing, and coordination, is tightly regulated by a cell cycle control feedback loop system. Cyclin – Cdk complexes are extremely important in these processes (Figure ). The loop consists of several significant stages and checkpoints: Gap 0 (G0) cell resting phase, Gap 1 (G1), first checkpoint, Synthesis (S), DNA replication phase, Gap 2 (G2) second checkpoint, and Mitosis (M), cell division stage.
A cancer cell is a cell that has decided to become immortal and to divide indefinitely. In order to evade well established suicide or senescence program cancer cell uses genetic material mutations thereby gaining the ability to grow out of control and spread into it’s surrounding destroying normal cells in the process. Genes that acquire mutations leading to cancer phenotype can either belong to the tumour suppressor family of genes responsible for keeping the cells in check and stopping them from becoming cancerous. Second family of genes that acquire mutations are oncogenes – tumour promoting genes responsible for cell proliferation with a potential of giving a raise to cancer cell. When the genes become mutated they either lose their ability to function or have their function compromised depending on the type of mutation.
Some cancers are genetic in nature i.e. involve a very specific type of genes with a very specialized functions such as BRCA1 and 2 mutations in breast cancer. Recently epigenetic nature of many other types of cancers was discovered.
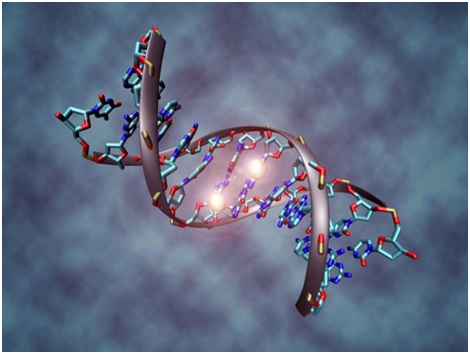
This is however where things get further complicated. Mutations can occur in genes involved in epigenetic regulation. However, hallmark of many cancers are epigenetic modifications related to epigenetic code which sits above our DNA and affects the way that our genes express themselves. Of all cancer specific epigenetic modifications characterized so far best understood and most extensively studied is DNA methylation.
DNA sequence consist of base pairs, abbreviated C, G, T and A. C, or cytosine nucleotide only ever base pairs with G base, or guanine nucleotide, by a phosphate whilst A adenosine pairs with corresponding T thymidine. When we speak of DNA methylation we essentially mean the process of tagging the C base of DNA with methyl groups. In this way our genes are chemically altered, but the genetic sequence itself isn't changed. Once a gene is chemically altered in this way it will be silenced or not expressed. Regions of the genome that are typically silenced in this way are long stretches of repetitive, retroviral, non-coding DNA regions which in the past we used to refer to as “junk DNA” but now days are only starting to appreciate its significance in health and disease. However, we are still far off from understanding the exact function and purpose of these genomic landscapes. Genomic regions which are typically methylation free are called gene promoters or CpG islands. Promoters are like a small switches or engines of a particular gene. This is where most of the crucial events for the gene happen. Decisions when to transcribe a gene, how much and for how long are made in these important regions.
They are very CG bases rich but almost always in an active genes they exclude methylation. Unless a gene needs to be silenced. Best studied example of gene silencing by methylation is X chromosome inactivation or so called imprinting. As you are probably aware male and female humans are genetically identical with the same number of chromosomes -23 pairs. Major difference is in their sex chromosomes males having XY and females XX sex chromosome. Y chromosome is fairly gene poor whilst X chromosome contains over 1000 genes essential for cellular development and viability. Females therefore carry two X chromosomes and are faced with potentially lethal dosage problem. This imbalance is corrected in females by inactivation of one of theirs X chromosomes through a mechanism of dosage compensation. One of X chromosomes is methylated and inactivated in females. Non coding RNA called Xist RNA helps in silencing by coating the chromosome.
So this is the state of the epigenome in health. In cancer however there is a complete breakdown at this level of regulation. There is a global state of hypo methylation with regions that should be methylated and silenced not being so any longer. There is also a state of hyper methylation, and this occurs in the promoter regions which are normally methylation free. In cancer, promoters of tumour suppressor genes become most frequent targets of silencing by DNA methylation. Consequence of this to the cell is detrimental. Cell is no longer kept in check and starts to proliferate abnormally. What causes the promoter of a gene which is normally in health methylation free to succumb to methylation and silencing is a subject of intense research.
Part of the answer probably lies with observed genetic mutations and loss of function of the family of enzymes called DNA methyltransferases (DNMTs) which are responsible for DNA methylation and the family of enzymes called TETs involved in demethylation but this is subject of ongoing research. Nobody is really entirely sure how does methylation process goes wrong. It is possible that the lifetime experience of tissue damage resulting from inflammation, environmental exposure or aging results in broken down epigenome. It has certainly been observed that the older people exhibit more epigenome damage than the younger ones. Furthermore diet is considered to be a contributing factor leading to DNA methylation alterations. Folate and Methionine pathways provide methyl groups required for DNA and histone methylation. We cannot synthesise these compound and the only way we can get them is through nutrition. Diet low in methionine and folate leads to DNA methylation level changes which have been associated with cancer. Certain agents such as arsenic and cadmium were shown to affect DNA methylation. Arsenic used in production of herbicides, pesticides and insecticides can lead to hypomethylation of ras oncogene an important cellular growth signalling oncogene. Cadmium chemical used in steel plating and plastics stabilizing was shown to induce global DNA hypomethylation through DNMT1 enzyme inactivation.
Attempts by the cells over the years to repair the damage may result in the loss of ability to perform this function efficiently, because the more frequently the repair is required the more likely it is that something will go wrong. It appears that following a certain number of division’s cellular epigenetic machinery starts making mistakes when tagging the genes.
Apart from the chemical changes to the DNA cancer epigenome is also characterized by significant changes to the so called “histone code”.
Here is one astonishing fact for you to consider. There is about 6 million base pairs of DNA in each cell which when stretched out is around 2 meters long. Human body on average contains 50 trillion cells with estimated 100 trillion meters of DNA. That is long enough to go from here to the Sun and back over 300 hundred times. So how do you pack a 2 meters long DNA string into the cell nucleus of roughly 10 micrometre in diameter? The answer lies in DNA condensation by coiling around specific proteins called histones (figure).

DNA is compacted into the tiny cell nucleus in more or less condensed form called heterochromatin (more tightly packed) or euchromatin (loosely packed). The more compact the DNA, the less likely it is for those particular genes to be expressed and Vice versa less compacted forms are usually regions of activity. Histone code is a set of chemical modifications added to the tail of the histones. The marks are like indexing cards tagging the regions of the genome which need to be expressed, silenced or poised. Some of the marks are found in more condensed regions marking inactive genes whilst others reside in open active area of the genome. Modifications to the tails of histone are extensively studied as well as the enzymes involved in laying down or removing particular marks. Aberrant histone modifications have oncogenic potential and are another hallmark of cancer as they also can affect gene activity.
DNA methylation and histone modifications are not only epigenetic components with the ability to affect the way chromatin is regulated. Long non coding, micro RNA and small interfering RNAs are additional level of transcriptional control that the cell has at its disposal. Micro RNA is aberrantly expressed in cancer cells and has the ability to affect the genes involved in cell cycle.
It is becoming clear that the path of cancer initiation and progression involves an interplay between genetics and epigenetics. Various scenarios should be considered as to which of the mechanisms are associated with cancer invitation and which with progression. In inherited cancer main initiator is a genetic mutation, with epigenetics being probably responsible for the disease progression. In other types of cancer possible scenario is that genetic mutations result in cellular transformation but its epigenetic changes that determine whether metastasis occurs. Finally, it should be considered that epigenetic process may be responsible for cancer imitation. For example epigenetic changes could prime the cells to promote cellular transformation with a subsequent DNA mutation occurring. In future articles we will be taking a closer look into most extensively studied cancer associated genes such as apoptosis regulating p53, INK/ARF tumour suppressor locus, BRCA1 and 2 and so on and genetic as well as epigenetic components of their governance.
Discovery of epigenetic nature of many cancer types has led to significant advances in the field of cancer biomarker discovery and cancer therapy. Human Epigenome Project (HEP) aims to integrate high throughput sequencing technology and huge datasets with epigenome sequencing. The end result it is hoped will give greater insight into epigenetic marks and their function in various cells and tissues and at various stages of disease state. The bigger the insight the greater the chance of development of specific tools to diagnose the patients early and determine the severity of their disease.
Therapeutic epigenetics is another area with a potential to benefit from this vibrant research. The very nature of epigenetic process lends itself to chemical intervention in a way not possible with genetic mutations. Radiation, chemotherapy and surgery are a first generation of cancer therapy. Next generation could well be epigenetic therapy. Two epigenetic drugs are already on the market Vidaza (azacitadine) DNA demethylating agent used to treat Acute Myelogenous leukaemia AML and myelodyspalstic syndrome MDS. Second is histone deactelyase inhibitor used to treat T cell lymphoma.
Discovery that epigenetic modifications are reversible in principle lead to great advances being made in development of epigenetically based prevention/intervention strategies. Factors such as diet modification and lifestyle intervention could be used to achieve a degree of epigenetic reprogramming to help maintain normal gene expression status and reverse tumorigenesis. Dietary compounds can influence cell epigenome and ultimately individual's risk of developing cancer. In future articles we will attempt to summarize most current findings from animal and human studies, on the capacity of nutrients to influence epigenetic processes. Targeting epigenetic pathways through diet-based intervention is promising area of research with great potential of translating these scientific findings into public health policies for the purpose of cancer prevention.